For several years now, we have talked frequently about the end of the age of antibiotics. As a result of over-prescribing and wide-spread use in the meat, poultry, and dairy industries–not to mention fish farming and the ability of pathogens to easily evolve around traditional man-made antipathogens–the standard arsenal of antibiotics is under severe stress.
About 700,000 globally die each year due to drug-resistant infections including tuberculosis, HIV, and malaria.1 Jim O’Neill. “Review on Antimicrobial Resistance.” AMR-Review. (Accessed 29 Oct 2017.) http://amr-review.org/home.html But experts predict that could rise to 10 million people a year by 2050 if no action is taken. In its recently released Antibacterial Agents in Clinical Development report, The World Health Organization (WHO) labeled this resistance “a global health emergency”, warning it could “seriously jeopardize” progress made in modern medicine.2 “ANTIBACTERIAL AGENTS IN CLINICAL DEVELOPMENT: An analysis of the antibacterial clinical development pipeline, including tuberculosis.” WHO. September 2017. http://apps.who.int/iris/bitstream/10665/258965/1/WHO-EMP-IAU-2017.11-eng.pdf In addition, if antibiotics continue to lose their effectiveness, then many common medical procedures – including intestinal surgery, C-sections, joint replacements, and even chemotherapy – could become too dangerous to perform.
About 700,000 globally die each year due to drug-resistant infections including tuberculosis, HIV, and malaria.
The WHO report also noted that, currently, there are few potential treatment options for those antibiotic-resistant infections–including drug-resistant tuberculosis which kills around 250,000 people each year.3 Karin Weyer, Dennis Falzon, Ernesto Jaramillo, et al. “DRUG-RESISTANT TUBERCULOSIS: WHAT IS THE SITUATION, WHAT ARE THE NEEDS TO ROLL IT BACK.” AMR Control. Jul 31, 2017 | 2017, Healthcare | 0. http://resistancecontrol.info/2017/drug-resistant-tuberculosis-what-is-the-situation-what-are-the-needs-to-roll-it-back-2/ There is also a lack of treatment options for bacteria groups including Acinetobacter and Enterobacteriaceae, such as Klebsiella and E.coli — which are commonly spread in hospitals and nursing homes and are often fatal.
So, what’s being done about it? According to the WHO report, as of May 2017, a total of 51 antibiotics (including combinations) were in the clinical pipeline. Unfortunately, most of the agents in the pipeline are modifications of existing antibiotic classes. As such, they offer only short-term solutions as they usually can’t overcome multiple existing resistance mechanisms and do not control the growing number of pan-resistant pathogens. In fact, out of the 51 antibiotics under development, only nine can be classified as innovative, possible long-term solutions.
With that in mind, let’s look at some of the innovations in the pipeline that just might hold some real, long-term hope.
It’s About Time
 I’ve been saying for several decades that the fundamental difference between manmade antibiotics and nature-based defenses against pathogens is that manmade antibiotics are one-trick ponies around which pathogens can easily evolve. Nature, on the other hand, whether it’s your own immune system or a natural antibiotic such as garlic, uses a multi-pronged approach that is far more difficult to work around. And still most researchers hunt desperately for the next one-trick pony–the next new antibiotic that will be able to function effectively for maybe a couple of years until it too is rendered useless. Fortunately, some researchers have finally decided to learn from nature rather than ignore it.
I’ve been saying for several decades that the fundamental difference between manmade antibiotics and nature-based defenses against pathogens is that manmade antibiotics are one-trick ponies around which pathogens can easily evolve. Nature, on the other hand, whether it’s your own immune system or a natural antibiotic such as garlic, uses a multi-pronged approach that is far more difficult to work around. And still most researchers hunt desperately for the next one-trick pony–the next new antibiotic that will be able to function effectively for maybe a couple of years until it too is rendered useless. Fortunately, some researchers have finally decided to learn from nature rather than ignore it.
University of Michigan researchers, for example, in collaboration with colleagues at Harvard University, have discovered a key difference between the way immune cells attack bacteria and the way antibiotics do. Where drugs typically attack a single process within bacteria, T cells attack a host of processes at the same time.4 Farokh Dotiwala et al. “Granzyme B Disrupts Central Metabolism and Protein Synthesis in Bacteria to Promote an Immune Cell Death ProGram.” Cell (2017). Volume 0 , Issue 0. http://www.cell.com/cell/fulltext/S0092-8674(17)31188-1
Killer T cells, formally known as cytotoxic lymphocytes, attack infected cells by producing the enzyme granzyme B. How this enzyme triggers death in bacteria has not been well understood until now. The researchers used proteomics–a technique that measures protein levels in a cell–and computer modeling to see granzyme B’s multipronged attack targeting multiple processes within the bacterial cell worked against different threats such as: E. coli, listeria, and tuberculosis.
As it turns out, the cytotoxic, granule-granzyme, proteases released by cytotoxic lymphocytes trigger oxidative bacterial death by:
- Disrupting electron transport
- Generating a superoxide anion
- And inactivating bacterial oxidative defenses
In addition, they also cause non-oxidative cell death because anaerobic bacteria are also killed. When it comes to anaerobic bacteria such as Escherichia coli, Listeria monocytogenes, and Mycobacteria tuberculosis, granzyme B cleaves a highly conserved set of proteins in all three bacteria, which function in vital biosynthetic and metabolic pathways that are critical for bacterial survival under diverse environmental conditions. Because killer cells use a multipronged strategy to target vital pathways, bacteria cannot easily become resistant to killer cell attack.
Sriram Chandrasekaran, one of the researchers, said, “When exposed to granzyme B, the bacteria were unable to develop resistance to the multipronged attack, even after exposure over multiple generations. This enzyme breaks down multiple proteins that are essential for the bacteria to survive. It’s essentially killing several birds with one stone.”
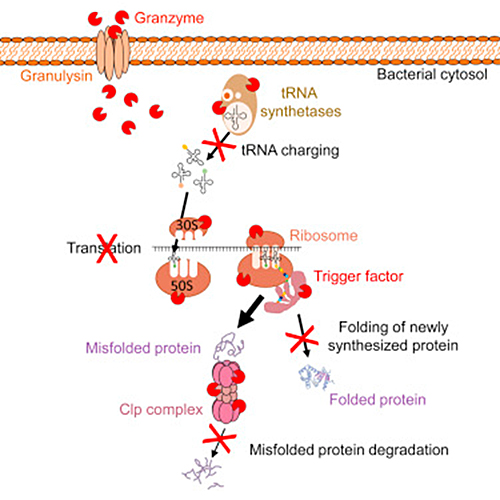
The possible applications of the new findings on T cells run the gamut from the creation of new medications to the repurposing of previously approved drugs in combination to fight infections by mimicking granzyme B.
Chandrasekaran’s team is now looking at how bacteria do eventually hide to avoid T-cell attacks.
Tweaking a Compromised Standard
Vancomycin is an antibiotic that is recommended intravenously as a treatment for complicated skin infections, bloodstream infections, endocarditis, bone and joint infections, and meningitis caused by methicillin-resistant S. aureus. Vancomycin belongs to a group of antibiotics called lipopeptides that work by compromising the cell walls of invading pathogens. Bacteria have an external cell wall that is reinforced by molecules called peptidoglycans. The cell wall protects the bacteria from your body’s normal environment in which the bacteria live. Vancomycin works by blocking the formation of these peptidoglycans. Specifically, vancomycin attaches itself to the walls of Gram-positive bacteria (built from linked amino acids D-Alanine-D-Alanine). Once attached, vancomycin simply prevents the D-Alanine amino acids from properly joining together, which weakens the cell wall–to the point the bacterial cell wall eventually crumbles, allowing the insides of the cell to flow out, thereby killing the cell. Note: the cell wall structure of Gram-negative bacteria is different, so vancomycin doesn’t work on them. Think E. coli, gonorrhea, cholera, syphilis, and Lyme disease.
For years, vancomycin was considered the option of last resort to treat many bacterial infections. Unfortunately, many Gram-positive bacteria, including enterococci (e.g., E faecalis and E faecium) and MRSA, have developed resistance to vancomycin–not really surprising when you consider how easily bacteria can develop resistance to one-trick ponies such as lipopeptides. The fact that it took several decades is the surprise.
Encouragingly, scientists have recently announced that they have been able to modify the drug in the lab so that it now has three distinct molecular moves to take out pathogens instead of just one.5 Akinori Okano, Nicholas A. Isley, and Dale L. Boger. “Peripheral modifications of [Ψ [CH2NH]Tpg4]vancomycin with added synergistic mechanisms of action provide durable and potent antibiotics. PNAS 2017 114 (26) E5052-E5061. http://www.pnas.org/content/114/26/E5052
- First, they tweaked vancomycin to attach to an additional part of the pathogen cell wall. Dual D-Ala-D-Ala/D-Ala-D-Lac binding inhibits transpeptidase AND cell wall biosynthesis. Or in plain English, this tweak totally bypasses current bacterial resistance to Vancomycin.
- They also added (4-chlorobiphenyl)methyl or CBP, which inhibits the transglycosylase enzyme, which is involved in bacterial cell wall construction.
- Finally, they added a quaternary ammonium salt to the structure to induce cell membrane permeability. Or to put it another way: to punch holes in the bacterial cell wall.
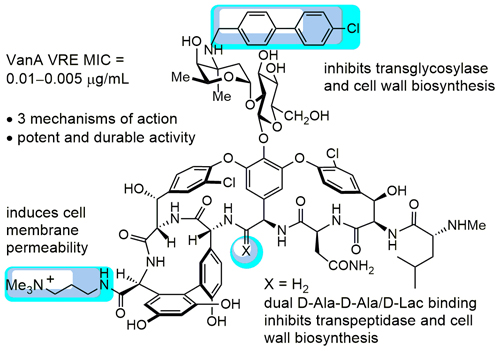
Three modifications to vancomycin’s structure (shown in blue) give it new “mechanisms of action” for fighting infection.
Thanks to the power of exponentiation, this makes new, triple-threat vancomycin 25,000-50,000 times deadlier than the old single-threat version. And it’s not only more effective, it’s also that much more impervious to bacteria developing resistance–something proven in lab experiments conducted as part of the study.
Unfortunately, as promising as this may be, it’s a long way from widespread human use. Even before it can be tested on humans in a clinical setting, it must go through animal testing and clinical trials to ensure safety and efficacy. Additionally, the scientists also need to develop methods for mass producing the altered vancomycin, as it currently requires about 30 steps. All in all, it will be several years, at best, before it is available.
As a side note, most antibiotics target Gram-positive bacteria, while the major challenge is to find new antibiotics against Gram-negative bacteria, which are identified as critical by WHO in its priority pathogens list for R&D of new antibiotics. Because of the complexity of the Gram-negative cell wall, discovery of novel antibiotics that can permeate this barrier and stay inside the bacterium is particularly challenging.
New, triple-threat vancomycin is 25,000-50,000 times deadlier than the old single-threat version.
Targeting Bacterial Enzymes
Instead of destroying a cell from the outside in, like vancomycin, some antibiotics bypass the cell-wall and block a cell’s ability to replicate from the inside out. Erythromycin, for example, works by binding to specific molecules — subunits — in a cell’s ribosome, destroying the cell’s ability to form the proteins it needs for cell growth. Sulfa antibiotics (sulfonamides) have been used to battle bacterial infections since the 1930s. They target specific chemical reactions within a cell — the metabolic pathways — by binding to an enzyme called dihydropteroate synthase (DHPS), which then blocks bacteria’s ability to synthesize dihydroazole acid, which the bacterium uses to process folate. When this type of bacterial cell stops being able to metabolize folate, it can no longer grow or multiply.
But bacteria have a defense against these types of antibiotics. They use enzymes known as beta-lactams to neutralize these types of antibiotics. One of these, β-lactamase, can potentially destroy any antibiotic that makes its way inside a cell.
But new research published in the Journal of Antimicrobial Chemotherapy6 Juan-Carlos Jiménez-Castellanos, Wan Ahmad Kamil Wan Nur Ismah, Yuiko Takebayashi, Matthew B Avison, et al. “Envelope proteome changes driven by RamA overproduction in Klebsiella pneumoniae that enhance acquired ß-lactam resistance.” Journal of Antimicrobial Chemotherapy, dkx345. http://academic.oup.com/jac/article-abstract/doi/10.1093/jac/dkx345/4364866?redirectedFrom=fulltext and Molecular Microbiology7 Karina Calvopiña,Philip Hinchliffe,Jürgen Brem,Matthew B. Avison, et al. “Structural/mechanistic insights into the efficacy of nonclassical ß-lactamase inhibitors against extensively drug resistant Stenotrophomonas maltophilia clinical isolates.” Molecular Biology. Volume 106, Issue 3 November 2017 Pages 492–504. http://onlinelibrary.wiley.com/doi/10.1111/mmi.13831/abstract indicates that targeting beta-lactams may offer the best chance of overcoming this type of antibiotic resistance. The researchers found that a combination of an enzyme inhibitor called avibactam and a β-lactam antibiotic called aztreonam could kill some of the most resistant bacteria known.
Dr. Matthew Avison, one of the researchers involved in the work, said: “Our bacteriology research has further demonstrated that beta-lactamases are the real ‘Achilles heel’ of antibiotic resistance in bacteria that kill thousands of people in the UK every year.8 “Scientific breakthrough may stop post-antibiotic apocalypse.” Angle News. 23 Oct 2017. (Accessed 28 Oct 2017.) http://www.anglenews.com/scientific-breakthrough-may-stop-post-antibiotic-apocalypse/
“Structural/mechanistic work on beta-lactamase enzymes … is helping to drive the discovery of wave after wave of beta-lactamase inhibitors, including the potentially game-changing bicyclic boronated class, shown to be effective in our research, and recently successful in Phase I clinical trials.
“This is an exciting time for researchers studying beta-lactamase inhibitors. At the risk of sounding like King Canute, it is the first time for a decade that there is some genuine positivity about our ability to turn back the rising tide of beta-lactam antibiotic resistance.”
The bottom line, as the researchers said, is that aiming for the beta-lactamase enzyme could eventually make it possible to reverse a “significant proportion” of antibiotic resistance.
Where Things Stand Today
 All the new research that we’ve just discussed is very exciting, but the reality, here on the ground, is that the incidence of antibiotic resistant infections and deaths is climbing steadily–and projected to continue to do so into the future. Some of the species in which resistance is of greatest public health concern are listed below.
All the new research that we’ve just discussed is very exciting, but the reality, here on the ground, is that the incidence of antibiotic resistant infections and deaths is climbing steadily–and projected to continue to do so into the future. Some of the species in which resistance is of greatest public health concern are listed below.
Bacteria – Community
- Escherichia coli
- Mybocaterium tuberculosis (cause of tuberculosis)
- Neisseria gonorrhoeae (cause of gonorrhea)
- Salmonella Typhi
- Staphylococcus aureus, including community-associated MRSA (Methicillin-Resistant S. aureus)
- Streptococcus pneumoniae
Bacteria – Hospitals
- Acinetobacter baumannii
- Enterococcus faecium and Enterococcus faecalis, including VRE (Vancomycin-resistant enterococci)
- Multidrug-resistant enteric pathogens, including Escherichia coli and Klebsiella pneumoniae producing ESBL and KPC enzymes
- Pseudomonas aeruginosa
- Staphylococcus aureus, including MRSA (Methicillin-Resistant S. aureus)
- Stenotrophomonas maltophilia
Bacteria – Zoonotic disease
- Campylobacter species
- Salmonella species
Fungi
- Candida albicans
Parasites
- Leishmania species
- Plasmodium species (cause of malaria)
- Trypanosoma species
Viruses
- Cytomyegalovirus
- Herpes simplex virus
- HIV
With that in mind, let’s look at a handful of these villains, as representative of the entire list, to see how they’re trending.
Tuberculosis
According to the World Health Organization:9 “Drug-resistant TB: global situation.” WHO (Accessed 29 Oct 2017.) http://www.who.int/tb/areas-of-work/drug-resistant-tb/global-situation/en/
- Drug resistance surveillance data show that an estimated 480,000 people developed MDR-TB in 2014 and 190,000 people died.
- Extensively drug-resistant TB (XDR-TB) has been reported by 105 countries in 2014. On average, an estimated 9.7% of people with MDR-TB have XDR-TB.
MRSA
About 85 percent of serious, invasive MRSA infections are health care-associated infections.10 Review By: Stanley J. Swierzewski, III, M.D. “MRSA Infection Incidence & Prevalence.” HealthCommunities.com. Published: 19 Feb 2008. Last Modified: 15 Sep 2015. (Accessed 29 Oct 2017.) http://www.healthcommunities.com/mrsa-infection/incidence.shtml That is, they are contracted while patients are in health-care facilities. The incidence of community-associated MRSA varies according to population and geographic location. In the United States, more than 94,000 people develop serious MRSA infection and about 19,000 die from infection each year.
According to the Centers for Disease Control and Prevention, each year in the United States, more than 290,000 hospitalized patients are infected with Staphylococcus aureus, and of these staph infections, approximately 126,000 are related to MRSA.
E. coli
A 2015 study out of Duke University School of Medicine found that the number of antibiotic-resistant E. coli infections seen in small, community hospitals has doubled in recent years.11 Joshua T. Thaden, Vance G. Fowler, Daniel J. Sexton, Deverick J. Anderson. Increasing Incidence of Extended-Spectrum ß-Lactamase-Producing Escherichia coli in Community Hospitals throughout the Southeastern United States. Infection Control & Hospital Epidemiology, 2015; 1. http://www.cambridge.org/core/journals/infection-control-and-hospital-epidemiology/article/increasing-incidence-of-extendedspectrum-lactamaseproducing-escherichia-coli-in-community-hospitals-throughout-the-southeastern-united-states/E84C1D82D5AC7F8396EE37EE663DBAC2/core-reader This finding should be cause for concern considering, according to the researchers, that more than half of all US patients are treated in these facilities.
These researchers also found a three-fold increase in antibiotic-resistant infections among community members who had limited exposure to health care settings. And these patients were found to acquire the “superbug” at an even faster rate than patients who had regular contact with hospitals.
What Can You Do About It?
 Yes, we may be entering the end of one age of antibiotics, but as we’ve discussed today, we’re about to enter another, even more promising age. Eventually, scientists are going to figure out how to once again regain the upper hand when it comes to battling pathogens–at least until those same pathogens once again figure out how to reassert their dominance over the next generation of pharmaceuticals. Fortunately, that’s likely to be far enough in the future to not be a concern for anyone reading this report. On the other hand, that new age of wonder drugs is not yet here. In fact, it’s several years off, and the end of the age of antibiotics as we know them is here now and only going to get progressively worse and deadlier before relief arrives. That leaves us with a problem, then. What do we do while waiting for salvation? The reality is that if you get hit with a drug resistant infection before that day arrives, you could be in for a world of hurt if forced to rely on the current arsenal of antibiotics, antivirals, and antifungals. And it’s only going to get worse over the next several years.
Yes, we may be entering the end of one age of antibiotics, but as we’ve discussed today, we’re about to enter another, even more promising age. Eventually, scientists are going to figure out how to once again regain the upper hand when it comes to battling pathogens–at least until those same pathogens once again figure out how to reassert their dominance over the next generation of pharmaceuticals. Fortunately, that’s likely to be far enough in the future to not be a concern for anyone reading this report. On the other hand, that new age of wonder drugs is not yet here. In fact, it’s several years off, and the end of the age of antibiotics as we know them is here now and only going to get progressively worse and deadlier before relief arrives. That leaves us with a problem, then. What do we do while waiting for salvation? The reality is that if you get hit with a drug resistant infection before that day arrives, you could be in for a world of hurt if forced to rely on the current arsenal of antibiotics, antivirals, and antifungals. And it’s only going to get worse over the next several years.
The bottom line is that to survive the years of transition you’re going to have to optimize your immune system, while at the same time keeping a supply of natural antipathogens in your medicine cabinet–for use when you need them.
Optimizing Your Immune System
If you want to maintain your health, increase your longevity, and minimize your risk of many illnesses, you must boost your immune system. It is important to build your immune system not just to protect yourself from cancer, age related diseases, or autoimmune disorders either. Your garden variety flu is responsible for some 250,000 to 500,000 deaths worldwide each year–about 36,000 in the US alone–each and every year. In fact, of the some 1,700 bacteria and viruses known to cause disease, historically, the flu virus has been the leading cause of death — only recently being surpassed by the HIV virus and the resulting AIDS.
And now, with drug resistant versions of these pathogens emerging and gaining strength, it makes more sense than ever to optimize your immune system to cut them off at the pass before they have a chance to establish themselves in your body.
Optimizing your immune system is your first line of defense against invading pathogens.
For more information on how to optimize your immune system–and why you need to–check out: /immunity/building-your-immune-system-ebola
Natural Antipathogens
The medical community’s arsenal of anti-pathogens is somewhat limited–90% is comprised of antibiotics (which, as we’ve discussed are under siege), 5% of antivirals, and 5% of cytokine therapies. (In truth, there are very few effective medical antivirals.) The discovery and use of all these drugs has represented one of the greatest achievements of medical science over the last hundred years. Unfortunately, thanks to arrogance and misuse, resistant strains of bacteria and viruses have proliferated like fear mongers in an Ebola epidemic. And in truth, this trend has proven almost impossible to stop because of the simple rules of evolution–at least for the next several years while we make our way through antibiotic purgatory.
If you come down with a drug resistant infection, your only hope may be to turn to Mother Nature for help.
So, until the day that those innovative miracle drugs that we talked about finally move out of the pipeline and into the mainstream, you’re on your own. If you come down with a drug resistant infection, your only hope may be to turn to Mother Nature for help. To learn how and why, check out: /colds-flus-infectious-diseases/using-anti-pathogens

References
| ↑1 | Jim O’Neill. “Review on Antimicrobial Resistance.” AMR-Review. (Accessed 29 Oct 2017.) http://amr-review.org/home.html |
|---|---|
| ↑2 | “ANTIBACTERIAL AGENTS IN CLINICAL DEVELOPMENT: An analysis of the antibacterial clinical development pipeline, including tuberculosis.” WHO. September 2017. http://apps.who.int/iris/bitstream/10665/258965/1/WHO-EMP-IAU-2017.11-eng.pdf |
| ↑3 | Karin Weyer, Dennis Falzon, Ernesto Jaramillo, et al. “DRUG-RESISTANT TUBERCULOSIS: WHAT IS THE SITUATION, WHAT ARE THE NEEDS TO ROLL IT BACK.” AMR Control. Jul 31, 2017 | 2017, Healthcare | 0. http://resistancecontrol.info/2017/drug-resistant-tuberculosis-what-is-the-situation-what-are-the-needs-to-roll-it-back-2/ |
| ↑4 | Farokh Dotiwala et al. “Granzyme B Disrupts Central Metabolism and Protein Synthesis in Bacteria to Promote an Immune Cell Death ProGram.” Cell (2017). Volume 0 , Issue 0. http://www.cell.com/cell/fulltext/S0092-8674(17)31188-1 |
| ↑5 | Akinori Okano, Nicholas A. Isley, and Dale L. Boger. “Peripheral modifications of [Ψ [CH2NH]Tpg4]vancomycin with added synergistic mechanisms of action provide durable and potent antibiotics. PNAS 2017 114 (26) E5052-E5061. http://www.pnas.org/content/114/26/E5052 |
| ↑6 | Juan-Carlos Jiménez-Castellanos, Wan Ahmad Kamil Wan Nur Ismah, Yuiko Takebayashi, Matthew B Avison, et al. “Envelope proteome changes driven by RamA overproduction in Klebsiella pneumoniae that enhance acquired ß-lactam resistance.” Journal of Antimicrobial Chemotherapy, dkx345. http://academic.oup.com/jac/article-abstract/doi/10.1093/jac/dkx345/4364866?redirectedFrom=fulltext |
| ↑7 | Karina Calvopiña,Philip Hinchliffe,Jürgen Brem,Matthew B. Avison, et al. “Structural/mechanistic insights into the efficacy of nonclassical ß-lactamase inhibitors against extensively drug resistant Stenotrophomonas maltophilia clinical isolates.” Molecular Biology. Volume 106, Issue 3 November 2017 Pages 492–504. http://onlinelibrary.wiley.com/doi/10.1111/mmi.13831/abstract |
| ↑8 | “Scientific breakthrough may stop post-antibiotic apocalypse.” Angle News. 23 Oct 2017. (Accessed 28 Oct 2017.) http://www.anglenews.com/scientific-breakthrough-may-stop-post-antibiotic-apocalypse/ |
| ↑9 | “Drug-resistant TB: global situation.” WHO (Accessed 29 Oct 2017.) http://www.who.int/tb/areas-of-work/drug-resistant-tb/global-situation/en/ |
| ↑10 | Review By: Stanley J. Swierzewski, III, M.D. “MRSA Infection Incidence & Prevalence.” HealthCommunities.com. Published: 19 Feb 2008. Last Modified: 15 Sep 2015. (Accessed 29 Oct 2017.) http://www.healthcommunities.com/mrsa-infection/incidence.shtml |
| ↑11 | Joshua T. Thaden, Vance G. Fowler, Daniel J. Sexton, Deverick J. Anderson. Increasing Incidence of Extended-Spectrum ß-Lactamase-Producing Escherichia coli in Community Hospitals throughout the Southeastern United States. Infection Control & Hospital Epidemiology, 2015; 1. http://www.cambridge.org/core/journals/infection-control-and-hospital-epidemiology/article/increasing-incidence-of-extendedspectrum-lactamaseproducing-escherichia-coli-in-community-hospitals-throughout-the-southeastern-united-states/E84C1D82D5AC7F8396EE37EE663DBAC2/core-reader |












What is the potential for
What is the potential for increased side effects with these new more powerful antibiotics? If they are 25,000 times more powerful then a much lower dose could be used and maybe that would result in fewer side effects. On the other hand there may a much greater die off of beneficial bacteria and that is already a great enough problem.
Very offended that you chose
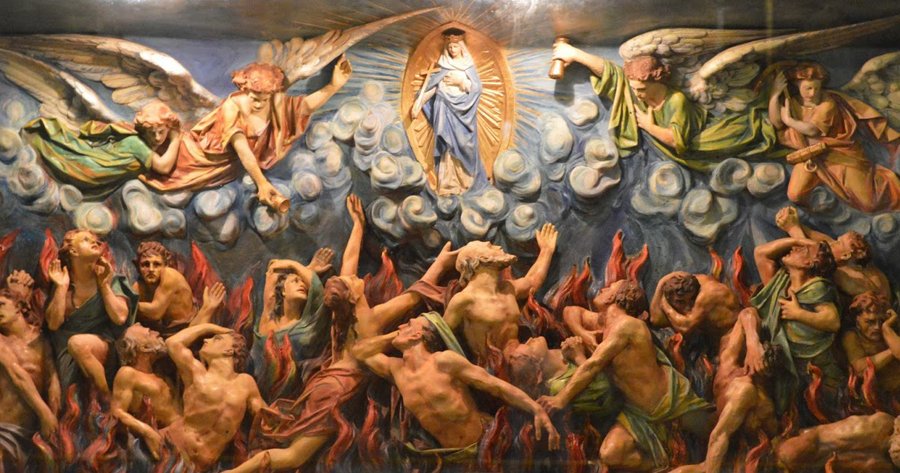
Very offended that you chose to compare “Purgatory” with an “Antibiotic Problem”. Do you even know what Purgatory is? It’s a place that Poor Souls (Catholics) who didn’t die in a State of Grace (no mortal sin) go to be purified before they can be in the presence of God in Heaven. They suffer terrible torments while waiting to be released. The month of November is the Month of Holy Souls where Catholics pray for and remember the Holy Souls in Purgatory.
What that has to do with your antibiotic article and the photo of Holy Souls suffering at the beginning of the article is beyond me.
Any suffering someone might go through while on Earth is of no comparison to the pain and torments that are endured while in Purgatory.
Don’t like the religious rant? Don’t include your offensive Purgatory reference in an article about antibiotics.
No comparison whatsoever, totally irrelevant and highly offensive to Catholics who take Purgatory seriously.
Sorry didn’t mean to offend.
Great points….In addition
Great points….In addition to boosting the immune system I would love for common practice to be these 3 simple things:
-Year round hand washing properly [20 seconds with soap) for ages 3 to 103
-sneezing and coughing into tissues not hands, desks, or seats ( drives me crazy)
-limiting sugar to daily intake of 70 grams or less (most people I know are shoving in much higher)
I also stopped taking really good food based multivitamins including zinc and vitamin c several years ago and noticed no difference in frequency/severity of colds (2-3 common colds a year). This leads me to believe that either the vitamins were not legit or my genetics are designed to run my immune system a certain way no matter what I do (though I don’t smoke or drink to really push things).
Thanks for your hard work!
At least I could not agree to
At least I could not agree to the proposition that there is nothing beyond antibiotics, mankind was never helpless nor will it ever be either in present or in future. For more than twenty-five years, I never used antibiotics, in fact I don’t believe in their efficacy, if it treats one then it develops ten other diseases. In fact, I never needed one, because homeopathic remedies are so effective that one’s anybody turns towards homeopathic remedies, he will never look back towards allopathic drugs.Blood transfusion, setting of broken bones, operations etc these are only techniques, which can be learnt by any system of medicines. Take simple echinacea, it can easily cure any kind of typhoid or par-typhoid,e-coli infection, incorrigible fever, septicaemia, and any other infection within seven days, mercurius solubilis in potentised form can easily cure syphilis, Curing MDR, XDR, TDR TBs are the easiest thing to treat in homeopathy. If timely homeo remedies are administered no TB patient will ever die, but no organisation not even the WHO wants to accept this breakthrough. With using certain homeo TB drugs we can prevent TB infection in the world, even these remedies are vigorously introduced then within a decade we can eradicate TB from the world, this is my belief. Unfortunately we do not find any takers, because there is a clique in the WHO and elsewhere, which people think that beyond their system of medicine, no other system is that effective. Unfortunately for mankind, for these people their system of medicine is more important than the welfare of humanity. Once I wrote a comprehensive letter to Chief Medical Officer UK on effectiveness of homeo drugs on treatment of TB, she simply informed me that millions of dollars have been pushed into research to find out new antibiotics, but she refrained from answering my offer. This is the tragedy! My claim is not hollow, more than hundreds of complicated TB cases I treated effectively administering only homeo drugs. Prostatitis is easily curable no matter whether cancerous or non cancerous in its early stage, benign cases are easily curable whatever be the stage, though it will take some time. Bone tumour is easily curable, same is the case with any thyroid cases whether hyper or hypo, no need to take medicine throughout life. Osteoporosis, Rheumatoid arthritis or any arthritis cases are easily amenable to momeo treatment no need for any steroid. Same is the case with leprosy or lupus, though they will take time and the patients must have faith. The world does not know at present nor will it believe what homeopathic remedies can do or undo to relieve the ailments of mankind. I cannot elaborate much here. But we have to remember the curability depends on, how much a patient has deteriorated his system by taking at random allopathic remedies earlier. Once chemo drugs are administered it will be difficult to provide relief to a cancer patient with homeo drugs later on, yes pain can be relieved If any day Nobel Prize in Medicine is given to any homeo practitioner then see people all over the world will jump after homeopathy, but that will never happen , because pharmaceutical companies will never allow this to happen in their own interest. This is the tragedy with humankind! … .
I like your posting about
I like your posting about homeopathic remedies. My father used them a lot and had a lot bottles of them for various ailments. He also was an amateur iridoligist. I know a bit about iridology too. What would help with paratyphoid (Salmonella) with pigeons and you also mentioned prostate problems in the beginning stage. I just recently read that colloidal silver may help with fungus in toenails. Any comments.
Just search our site for
Just search our site for articles we have written about your questions, thanks!